Partial Recognition Score
For an oligonucleotide frequency matrix FL-m+1,k={fij} of the length L and an arbitrary DNA sequence S=s1...si...sL of the same length L, the simplest recognition Score is calculated:
![]()
Formula (2) calculates the Score# value, the scale range of which is increasing with the growth of the site length L, and decreasing with the growth of the size k of the oligonucleotide alphabet {E1, ..., Ej, ..., Ek}. According to Zadeh's fuzzy sets (Zadeh, 1965), the Score values calculated by the formula (2) were transformed into the normalized partial recognition scale:
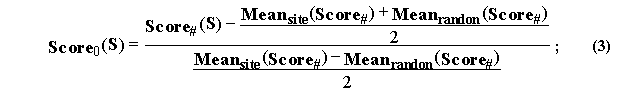
where the partial recognition’s rule is as follows:
IF {Score0(S)>0} THEN {S is this site}, OTHERWISE {S is not this site}.
Formula (3) gives the normalized Score0(S), the mean value of which averaged over all the known sequences of the site under study equals to “1”, whereas the mean value averaged over the random DNA sequences equals to “–1”. This Score0 scale is common for all the oligonucleotide frequency matrices compiled by the MATRIX database for any functional DNA site expressed within any alphabet.